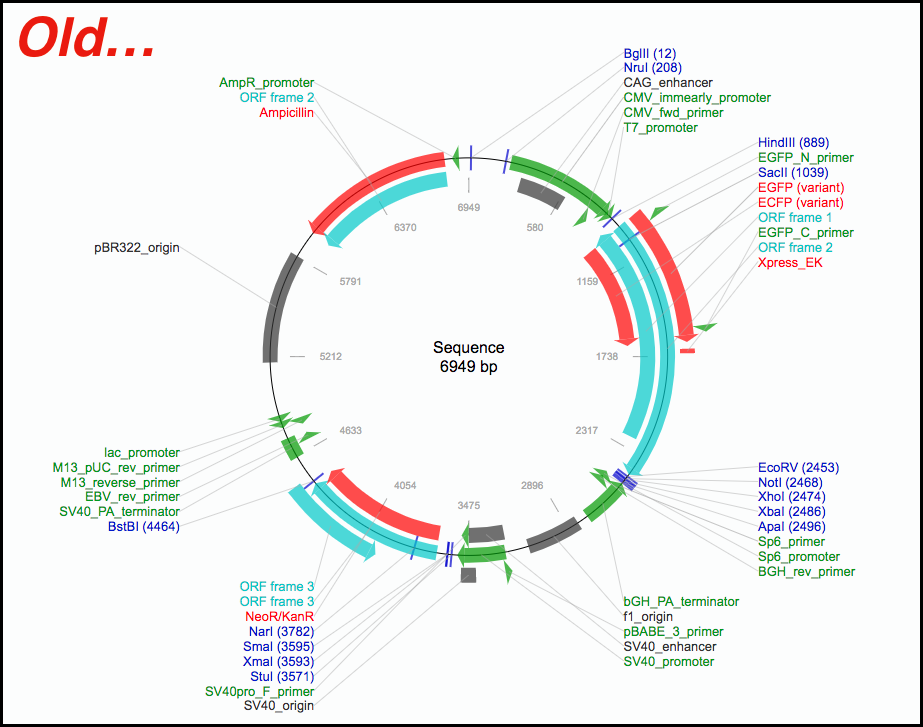
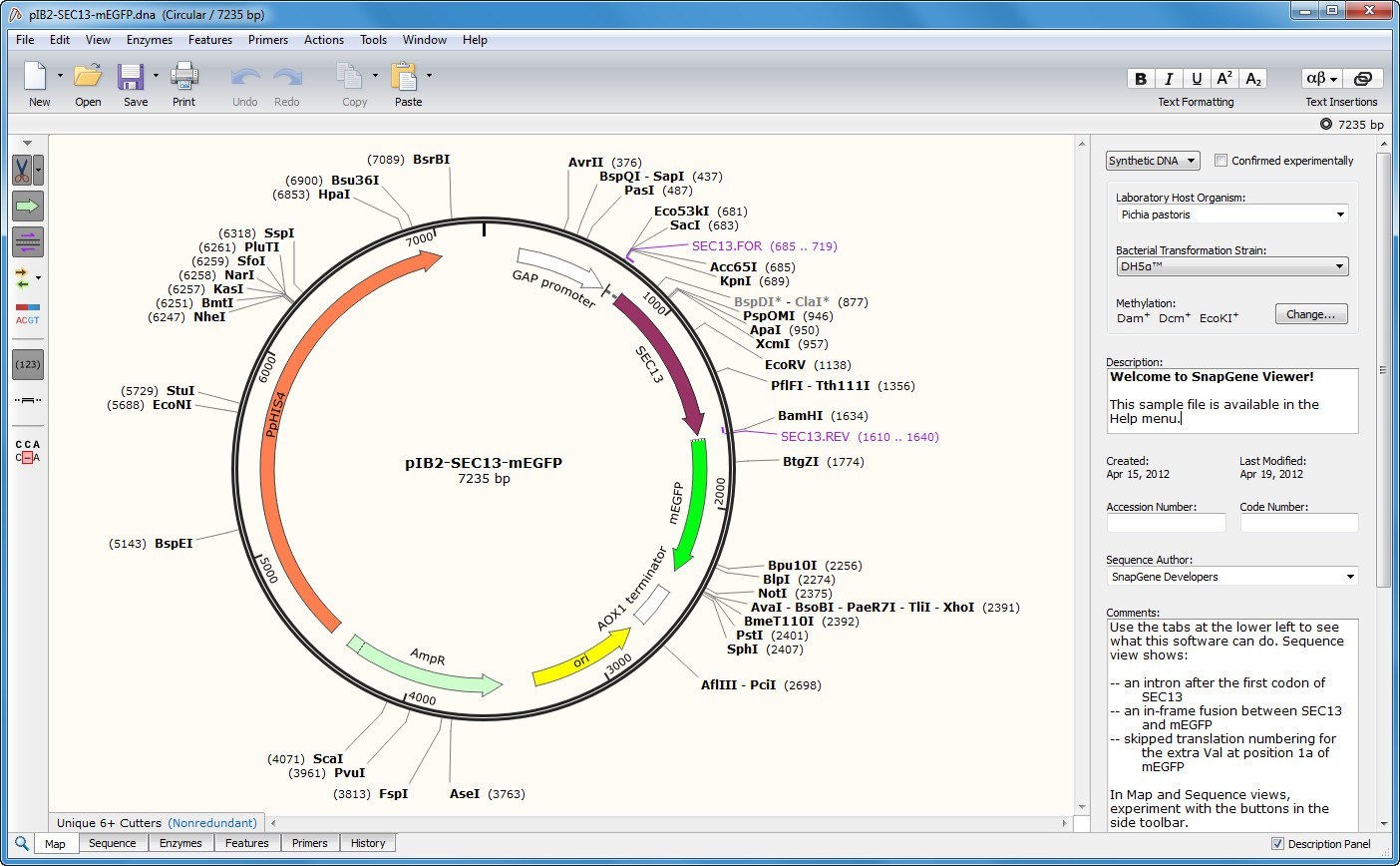
Analyze your plasmid sequenceįor more immediate, high-level analysis, you can click directly on the “Analyze Sequence” button to pull up an interactive plasmid map powered by SnapGene and gain access to additional tooltip information and sequence analysis tabs including: If the depositor provided a custom annotated GenBank file, this can be found on the main plasmid page under the "Resource Information" section. You can then analyze the sequence in your sequence viewer of choice. On this page you can download an annotated GenBank File or a SnapGene file for the sequences listed. Click on the “View all sequences” button to find all available sequence information from Addgene depositors and our own sequencing results: Once you’ve identified a plasmid that you want to learn more about, updates to our sequence analyzer will allow you to dive into the details. This will make it easier for you to share plasmid information with colleagues, or explain plasmid features to a lab member when you’re not at your computer. With this more functional display, you can start thinking about your next cloning experiment early on.īeyond these simple but powerful display improvements, when you click on any plasmid map, you also have the ability to to download the static image file (PNG, see above) for your own reference or to paste directly into your notebook. The list of enzymes that can be detected is 5x greater than our previous mapping software and now includes Type IIS restriction enzymes that are commonly used for Golden Gate cloning or for CRISPR gRNA cloning.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
